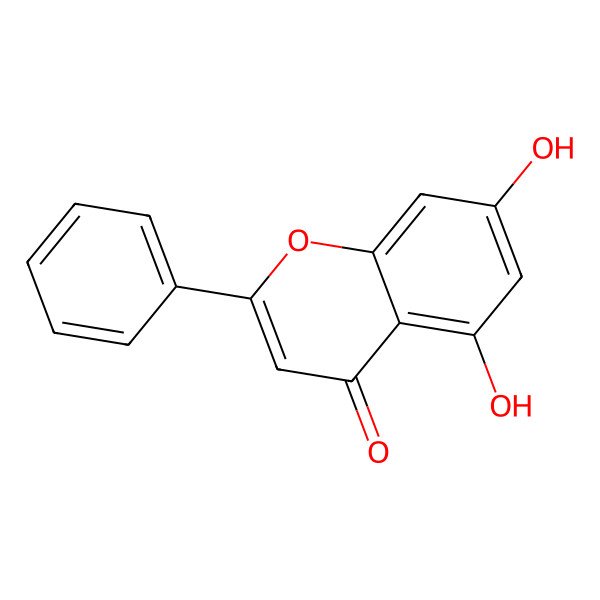
Chrysin
| Internal ID | 917579ab-a989-4ad9-9614-01814d58575f |
| Taxonomy | Phenylpropanoids and polyketides > Flavonoids > Flavones |
| IUPAC Name | 5,7-dihydroxy-2-phenylchromen-4-one |
| SMILES (Canonical) | C1=CC=C(C=C1)C2=CC(=O)C3=C(C=C(C=C3O2)O)O |
| SMILES (Isomeric) | C1=CC=C(C=C1)C2=CC(=O)C3=C(C=C(C=C3O2)O)O |
| InChI | InChI=1S/C15H10O4/c16-10-6-11(17)15-12(18)8-13(19-14(15)7-10)9-4-2-1-3-5-9/h1-8,16-17H |
| InChI Key | RTIXKCRFFJGDFG-UHFFFAOYSA-N |
| Popularity | 2,793 references in papers |
| Molecular Formula | C15H10O4 |
| Molecular Weight | 254.24 g/mol |
| Exact Mass | 254.05790880 g/mol |
| Topological Polar Surface Area (TPSA) | 66.80 Ų |
| XlogP | 2.10 |
| 480-40-0 |
| 5,7-Dihydroxyflavone |
| 5,7-Dihydroxy-2-phenyl-4H-chromen-4-one |
| Chrysine |
| 5,7-dihydroxy-2-phenylchromen-4-one |
| Crysin |
| 4H-1-Benzopyran-4-one, 5,7-dihydroxy-2-phenyl- |
| NSC-407436 |
| FLAVONE, 5,7-DIHYDROXY- |
| NSC 407436 |
| There are more than 10 synonyms. If you wish to see them all click here. |
| Target | Value | Probability (raw) | Probability (%) |
|---|---|---|---|
| No predicted properties yet! | |||
Proven Targets:
| CHEMBL ID | UniProt ID | Name | Min activity | Assay type | Source |
|---|---|---|---|---|---|
| CHEMBL1293255 | P15428 | 15-hydroxyprostaglandin dehydrogenase [NAD+] |
3981.1 nM 17782.8 nM |
Potency Potency |
via CMAUP
via CMAUP |
| CHEMBL3577 | P00352 | Aldehyde dehydrogenase 1A1 |
1584.9 nM 15848.9 nM 8912.5 nM 17782.8 nM |
Potency Potency Potency Potency |
via CMAUP
via CMAUP via CMAUP via CMAUP |
| CHEMBL1900 | P15121 | Aldose reductase |
8500 nM 7780 nM |
IC50 IC50 |
PMID: 18165015
PMID: 21256748 |
| CHEMBL5393 | Q9UNQ0 | ATP-binding cassette sub-family G member 2 |
2600 nM 1500 nM |
IC50 IC50 |
PMID: 21354800
PMID: 21354800 |
| CHEMBL261 | P00915 | Carbonic anhydrase I |
5490 nM 5420 nM |
Ki Ki |
PMID: 22487176
PMID: 22487176 |
| CHEMBL205 | P00918 | Carbonic anhydrase II |
4380 nM 4310 nM |
Ki Ki |
PMID: 22487176
PMID: 22487176 |
| CHEMBL3729 | P22748 | Carbonic anhydrase IV |
537.75 nM 537.75 nM |
Ki Ki |
via Super-PRED
PMID: 26498393 |
| CHEMBL3594 | Q16790 | Carbonic anhydrase IX |
2650 nM 2710 nM |
Ki Ki |
PMID: 22487176
PMID: 22487176 |
| CHEMBL2326 | P43166 | Carbonic anhydrase VII |
171.41 nM 171.41 nM |
Ki Ki |
PMID: 26498393
via Super-PRED |
| CHEMBL3242 | O43570 | Carbonic anhydrase XII |
8960 nM 34.7 nM 9070 nM |
Ki Ki Ki |
PMID: 22487176
via Super-PRED PMID: 22487176 |
| CHEMBL5586 | P16152 | Carbonyl reductase [NADPH] 1 |
670 nM 110 nM |
IC50 Ki |
PMID: 19097799
via Super-PRED |
| CHEMBL4096 | P04637 | Cellular tumor antigen p53 |
25118.9 nM 25118.9 nM |
Potency Potency |
via CMAUP
via CMAUP |
| CHEMBL2508 | Q00534 | Cyclin-dependent kinase 6 |
6000 nM 6000 nM |
IC50 IC50 |
DOI: 10.1007/s00044-009-9233-5
PMID: 15689157 |
| CHEMBL230 | P35354 | Cyclooxygenase-2 |
25500 nM |
IC50 |
PMID: 24332492
|
| CHEMBL1978 | P11511 | Cytochrome P450 19A1 |
1 nM 1 nM 1500 nM 500 nM 800 nM |
IC50 IC50 IC50 IC50 IC50 |
PMID: 20413308
via Super-PRED PMID: 22444875 PMID: 26301554 DOI: 10.1039/C3MD00166K |
| CHEMBL2231 | P04798 | Cytochrome P450 1A1 |
1290 nM 153 nM |
IC50 IC50 |
PMID: 23600958
PMID: 20696580 |
| CHEMBL3356 | P05177 | Cytochrome P450 1A2 |
84 nM 54 nM |
IC50 IC50 |
PMID: 20696580
PMID: 20832301 |
| CHEMBL4878 | Q16678 | Cytochrome P450 1B1 |
24 nM 16 nM |
IC50 Ki |
PMID: 20696580
via Super-PRED |
| CHEMBL3622 | P33261 | Cytochrome P450 2C19 |
794.3 nM |
Potency |
via CMAUP
|
| CHEMBL3397 | P11712 | Cytochrome P450 2C9 |
5225 nM |
IC50 |
PMID: 20832301
|
| CHEMBL289 | P10635 | Cytochrome P450 2D6 |
12589.25 nM |
AC50 |
via CMAUP
|
| CHEMBL340 | P08684 | Cytochrome P450 3A4 |
2511.9 nM 3162.3 nM 2511.9 nM 3162.3 nM |
Potency Potency Potency Potency |
via CMAUP
via CMAUP via CMAUP via CMAUP |
| CHEMBL4159 | Q99714 | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein |
10000 nM |
Potency |
via CMAUP
|
| CHEMBL4261 | Q16665 | Hypoxia-inducible factor 1 alpha |
6309.6 nM 6309.6 nM |
Potency Potency |
via CMAUP
via CMAUP |
| CHEMBL1293226 | B2RXH2 | Lysine-specific demethylase 4D-like |
31622.8 nM |
Potency |
via CMAUP
|
| CHEMBL4040 | P28482 | MAP kinase ERK2 |
31622.8 nM |
Potency |
via CMAUP
|
| CHEMBL1293224 | P10636 | Microtubule-associated protein tau |
15848.9 nM 35481.3 nM 35481.3 nM 5623.4 nM 11220.2 nM |
Potency Potency Potency Potency Potency |
via CMAUP
via CMAUP via CMAUP via CMAUP via CMAUP |
| CHEMBL1293232 | Q16637 | Survival motor neuron protein |
3981.1 nM 14125.4 nM 28183.8 nM |
Potency Potency Potency |
via CMAUP
via CMAUP via CMAUP |
| CHEMBL1929 | P47989 | Xanthine dehydrogenase |
840 nM |
IC50 |
PMID: 9461655
|
Predicted Targets (via Super-PRED):
| CHEMBL ID | UniProt ID | Name | Probability | Model accuracy |
|---|---|---|---|---|
| CHEMBL5619 | P27695 | DNA-(apurinic or apyrimidinic site) lyase | 98.63% | 91.11% |
| CHEMBL2581 | P07339 | Cathepsin D | 94.96% | 98.95% |
| CHEMBL3108638 | O15164 | Transcription intermediary factor 1-alpha | 94.39% | 95.56% |
| CHEMBL1806 | P11388 | DNA topoisomerase II alpha | 92.60% | 89.00% |
| CHEMBL3194 | P02766 | Transthyretin | 90.23% | 90.71% |
| CHEMBL1951 | P21397 | Monoamine oxidase A | 89.67% | 91.49% |
| CHEMBL1860 | P10827 | Thyroid hormone receptor alpha | 89.05% | 99.15% |
| CHEMBL1293249 | Q13887 | Kruppel-like factor 5 | 88.39% | 86.33% |
| CHEMBL3192 | Q9BY41 | Histone deacetylase 8 | 87.48% | 93.99% |
| CHEMBL3038477 | P67870 | Casein kinase II alpha/beta | 87.45% | 99.23% |
| CHEMBL2035 | P08912 | Muscarinic acetylcholine receptor M5 | 85.30% | 94.62% |
| CHEMBL2635 | P51452 | Dual specificity protein phosphatase 3 | 84.34% | 94.00% |
| CHEMBL3091268 | Q92753 | Nuclear receptor ROR-beta | 83.40% | 95.50% |
| CHEMBL3401 | O75469 | Pregnane X receptor | 81.50% | 94.73% |
| CHEMBL5284 | Q96RR4 | CaM-kinase kinase beta | 80.45% | 89.23% |
Below are displayed all the plants proven (via scientific papers) to contain this
compound!
To see more specific details click the taxa you are interested in.
To see more specific details click the taxa you are interested in.