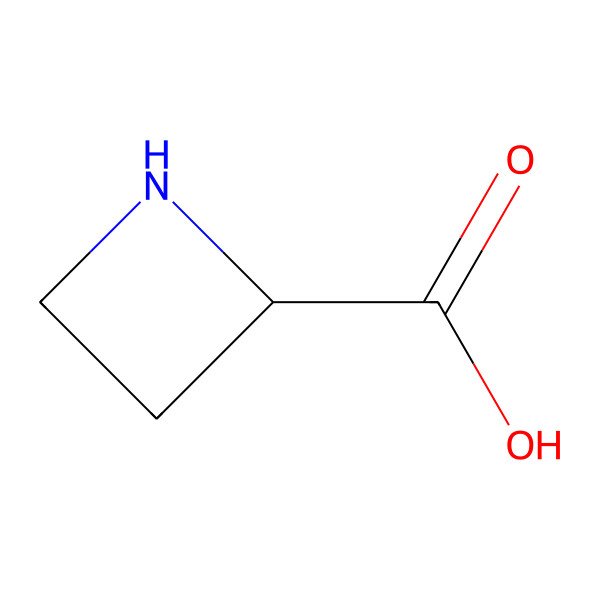
| L-Azetidine-2-carboxylic acid |
| (S)-Azetidine-2-carboxylic acid |
| (2S)-azetidine-2-carboxylic acid |
| (S)-(-)-2-Azetidinecarboxylic acid |
| (S)-2-Azetidinecarboxylic acid |
| Azetidyl-2-carboxylic acid |
| 2-Azetidinecarboxylic acid, (S)- |
| 2-Azetidinecarboxylic acid, (2S)- |
| (S)-(-)-Azetidine-2-carboxylic acid |
| L-Azetidine 2-carboxylic acid |
| Azetidine-2-carboxylic acid, L- |
| CHEBI:6198 |
| (L)-AZETIDINE-2-CARBOXYLIC ACID |
| Azetidinecarboxylic acid |
| 2-Azetidinecarboxylic acid, L- |
| (S)-L-Azetidine-2-carboxylic acid |
| MFCD00005166 |
| (S)-(-)-2-azetidine carboxylic acid |
| 5GZ3E0L9ZU |
| L-2-Azetidinecarboxylic acid |
| Acide L-azetidine-2-carboxylic |
| CHEMBL1165239 |
| L-azetidine-2-carboxylate |
| azetidine-2-carboxylate |
| HSDB 3465 |
| (2S)azetidine-2-carboxylic acid |
| AzeOH |
| 02A |
| EINECS 218-362-5 |
| L-AzeOH |
| L-Trimethyleneimine-2-carboxylic Acid |
| (S)-(-) Azetidine-2-carboxylic acid |
| Acide L-azetidine-2-carboxylic [French] |
| ST059592 |
| H-Aze(2)-OH |
| UNII-5GZ3E0L9ZU |
| L-azetidine-2caboxylic acid |
| (S)-Azetidin-2-carbonsaure |
| L-azetidine-2carboxylic acid |
| Lopac0_000023 |
| SCHEMBL20296 |
| (S)-2-azetidinecarboxyic acid |
| GTPL4686 |
| (S)-Azetidine-2-carboxylicacid |
| 2-(S)-azetidinecarboxylic acid |
| azetidine-2(S)-carboxylic acid |
| L--Azetidine-2-carboxylic acid |
| DTXSID0044020 |
| (S)-2-azetidine carboxylic acid |
| azetidine-2-(S)-carboxylic acid |
| HMS3260E07 |
| Tox21_500023 |
| (-)-AZETIDINECARBOXYLIC ACID |
| BDBM50357225 |
| AKOS005254687 |
| AKOS006239010 |
| (S)-(-)-2-azetidine-carboxylic acid |
| AC-5699 |
| CCG-204119 |
| HY-W050044 |
| LP00023 |
| SDCCGSBI-0050012.P002 |
| L-Azetidine-2-carboxylic acid, >=99% |
| NCGC00093546-01 |
| NCGC00093546-02 |
| NCGC00093546-03 |
| NCGC00093546-04 |
| NCGC00260708-01 |
| DS-16309 |
| (S)-2-Azetidinecarboxylic acid;H-Aze-OH |
| AZETIDYL-2-CARBOXYLIC ACID [HSDB] |
| A1043 |
| A4601 |
| AM20090185 |
| CS-0031389 |
| EU-0100023 |
| EN300-85821 |
| A 0760 |
| C08267 |
| Q793715 |
| SR-01000075656 |
| SR-01000075656-1 |
| W-107562 |
| azetidine-2-carboxylic acid;(S)-Azetidine-2-carboxylic acid |
| L-Azetidine-2-carboxylic acid is known as a proline analog. |
|
There are more than 10 synonyms. If you wish to see them all click here.
|