| 304-21-2 |
| Dihydroharmine |
| Harmidine |
| Armalin |
| Harmalol methyl ether |
| O-Methylharmalol |
| 3,4-Dihydroharmine |
| Harmine, dihydro- |
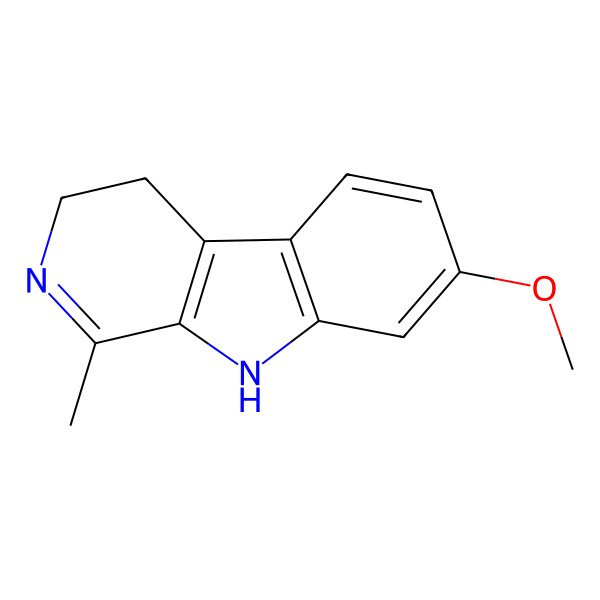
| 3H-Pyrido[3,4-b]indole, 4,9-dihydro-7-methoxy-1-methyl- |
| 7-methoxy-1-methyl-4,9-dihydro-3H-pyrido[3,4-b]indole |
| harmalin |
| NSC 407285 |
| 1-Methyl-7-methoxy-3,4-dihydro-beta-carboline |
| 3H-Pyrido(3,4-b)indole, 4,9-dihydro-7-methoxy-1-methyl- |
| 7-Methoxy-1-methyl-4,9-dihydro-3H-beta-carboline |
| 4,9-Dihydro-7-methoxy-1-methyl-3H-pyrido(3,4-b)indole |
| 7-methoxy-1-methyl-3,4-dihydro-2H-pyrido[3,4-b]indole |
| CN58I4TOET |
| 3,4-Dihydro-7-methoxy-1-methyl-9-pyrid(3,4-b)indole |
| 3,4-Dihydro-7-methoxy-1-methyl-9-pyrido(3,4-b)indole |
| MLS000028746 |
| 4,9-Dihydro-7-methoxy-1-methyl-3H-pyrido[3,4-b]indole |
| CHEMBL340807 |
| CHEBI:28172 |
| TNP00101 |
| NSC-407285 |
| SMR000058221 |
| 7-methoxy-1-methyl-4,9-dihydro-3H-b-carboline |
| 1-Methyl-7-methoxy-3,4-dihydro-.beta.-carboline |
| SR-01000721899 |
| EINECS 206-152-6 |
| UNII-CN58I4TOET |
| 3,4-Dihydro-7-methoxy-1-methyl-9-pyrid[3,4-b]indole |
| BRN 0207310 |
| Dihydro-Harmine |
| 7-METHOXY-1-METHYL-4,9-DIHYDRO-3H-PYRIDO(3,4-B)INDOLE |
| HSDB 7645 |
| MFCD00004955 |
| 3,4-b]indole |
| 3, 4-Dihydroharmine |
| Spectrum_000301 |
| Opera_ID_665 |
| HARMALINE [MI] |
| HARMALINE [HSDB] |
| Prestwick0_000610 |
| Prestwick1_000610 |
| Prestwick2_000610 |
| Prestwick3_000610 |
| Spectrum2_000411 |
| Spectrum3_000720 |
| Spectrum4_000824 |
| Spectrum5_001452 |
| 7-methoxy-1-methyl-3,4-dihydrobeta-carboline |
| HARMALINE [MART.] |
| 1-Methyl-7-methoxy-3,4-dihydro- beta-carboline |
| Oprea1_112644 |
| BSPBio_000520 |
| BSPBio_002539 |
| KBioGR_001367 |
| KBioSS_000781 |
| MLS001148231 |
| BIDD:ER0444 |
| DivK1c_000950 |
| SCHEMBL199260 |
| SPECTRUM1500864 |
| SPBio_000362 |
| SPBio_002739 |
| BPBio1_000572 |
| MEGxp0_001874 |
| SCHEMBL3862598 |
| DTXSID8041038 |
| ACon1_000019 |
| CHEBI:95328 |
| HMS502P12 |
| KBio1_000950 |
| KBio2_000781 |
| KBio2_003349 |
| KBio2_005917 |
| KBio3_001759 |
| NINDS_000950 |
| HMS1921K22 |
| HMS2233L23 |
| HMS3371G17 |
| HMS3885P22 |
| BDBM50029799 |
| CCG-38677 |
| NSC407285 |
| NSC789037 |
| s4776 |
| AKOS015904564 |
| AKOS026750619 |
| DB13875 |
| KS-5325 |
| NSC-789037 |
| SDCCGMLS-0066716.P001 |
| IDI1_000950 |
| NCGC00017221-01 |
| NCGC00017221-02 |
| NCGC00017221-03 |
| NCGC00017221-04 |
| NCGC00017221-05 |
| NCGC00017221-06 |
| NCGC00017221-07 |
| NCGC00094867-01 |
| NCGC00094867-02 |
| NCGC00094867-03 |
| NCGC00094867-04 |
| NCGC00094867-05 |
| NCGC00094867-06 |
| AC-34436 |
| LS-14204 |
| PD056221 |
| WLN: T B656 DM HM CHJ F1 KO1 |
| HY-107828 |
| 3,4-Dihydro-7-methoxy-1-methyl-b-carboline |
| CS-0030704 |
| FT-0626855 |
| H1237 |
| 3,4-Dihydro-7-methoxy-1-methyl-beta-carboline |
| C06536 |
| H-1100 |
| H-1102 |
| 1-Methyl-7-methoxy-3,4-dihydro-beta -carboline |
| 1-Methyl-7-methoxy-3, 4-dihydro-beta -carboline |
| 3H-Pyrido[3, 4,9-dihydro-7-methoxy-1-methyl- |
| 7-Methoxy-1-methyl-4,9-dihydro-3H-?-carboline |
| A820367 |
| Q135270 |
| 7-Methoxy-1-methyl-2,9-dihydro-1H-beta-carboline |
| 7-methoxy-1-methyl-3,4-dihydro-2h-beta-carboline |
| A1-00783 |
| SR-01000721899-4 |
| SR-01000721899-5 |
| 7-Methoxy-1-methyl-4,9-dihydro-3H-beta-carboline # |
| BRD-K91317041-001-03-8 |
| BRD-K91317041-310-03-3 |
| 3,4-Dihydro-7-methoxy-1-methyl-9H-pyrid[3,4-b]indole |
| 3,4-Dihydro-7-methoxy-1-methyl-9H-pyrido[3,4-b]indole |
|
There are more than 10 synonyms. If you wish to see them all click here.
|