Dioncophylline C
| Internal ID | bb282b25-8d4f-4eea-a668-843ae542cd1f |
| Taxonomy | Organoheterocyclic compounds > Isoquinolines and derivatives > Naphthylisoquinolines |
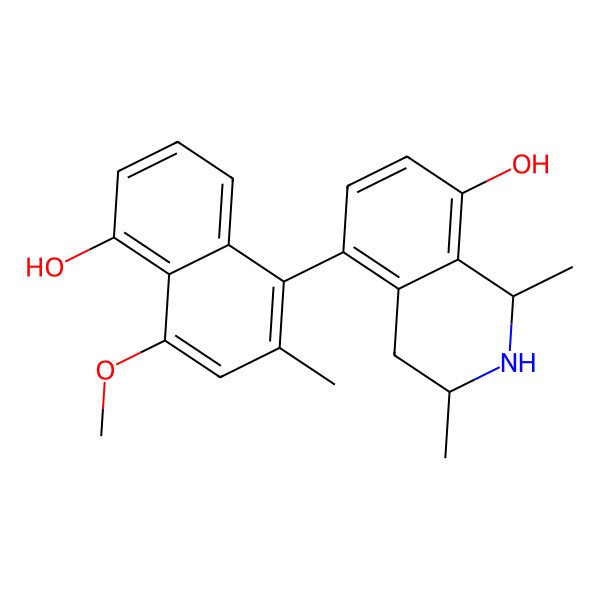
| IUPAC Name | (1R,3R)-5-(5-hydroxy-4-methoxy-2-methylnaphthalen-1-yl)-1,3-dimethyl-1,2,3,4-tetrahydroisoquinolin-8-ol |
| SMILES (Canonical) | CC1CC2=C(C=CC(=C2C(N1)C)O)C3=C4C=CC=C(C4=C(C=C3C)OC)O |
| SMILES (Isomeric) | C[C@@H]1CC2=C(C=CC(=C2[C@H](N1)C)O)C3=C4C=CC=C(C4=C(C=C3C)OC)O |
| InChI | InChI=1S/C23H25NO3/c1-12-10-20(27-4)23-16(6-5-7-18(23)25)21(12)15-8-9-19(26)22-14(3)24-13(2)11-17(15)22/h5-10,13-14,24-26H,11H2,1-4H3/t13-,14-/m1/s1 |
| InChI Key | NALOMJPIDNQZKW-ZIAGYGMSSA-N |
| Popularity | 25 references in papers |
| Molecular Formula | C23H25NO3 |
| Molecular Weight | 363.40 g/mol |
| Exact Mass | 363.18344366 g/mol |
| Topological Polar Surface Area (TPSA) | 61.70 Ų |
| XlogP | 4.70 |
| CHEBI:31505 |
| (1R,3R)-5-(5-hydroxy-4-methoxy-2-methylnaphthalen-1-yl)-1,3-dimethyl-1,2,3,4-tetrahydroisoquinolin-8-ol |
| CHEMBL375542 |
| Q27104194 |
| Target | Value | Probability (raw) | Probability (%) |
|---|---|---|---|
| No predicted properties yet! | |||
Proven Targets:
| CHEMBL ID | UniProt ID | Name | Min activity | Assay type | Source |
|---|---|---|---|---|---|
| No proven targets yet! | |||||
Predicted Targets (via Super-PRED):
| CHEMBL ID | UniProt ID | Name | Probability | Model accuracy |
|---|---|---|---|---|
| CHEMBL5619 | P27695 | DNA-(apurinic or apyrimidinic site) lyase | 99.03% | 91.11% |
| CHEMBL3108638 | O15164 | Transcription intermediary factor 1-alpha | 96.56% | 95.56% |
| CHEMBL1951 | P21397 | Monoamine oxidase A | 96.51% | 91.49% |
| CHEMBL2581 | P07339 | Cathepsin D | 96.07% | 98.95% |
| CHEMBL2535 | P11166 | Glucose transporter | 94.71% | 98.75% |
| CHEMBL1860 | P10827 | Thyroid hormone receptor alpha | 93.49% | 99.15% |
| CHEMBL241 | Q14432 | Phosphodiesterase 3A | 92.75% | 92.94% |
| CHEMBL1907 | P15144 | Aminopeptidase N | 92.69% | 93.31% |
| CHEMBL3251 | P19838 | Nuclear factor NF-kappa-B p105 subunit | 91.48% | 96.09% |
| CHEMBL1806 | P11388 | DNA topoisomerase II alpha | 91.41% | 89.00% |
| CHEMBL2041 | P07949 | Tyrosine-protein kinase receptor RET | 91.26% | 91.79% |
| CHEMBL5608 | Q16288 | NT-3 growth factor receptor | 90.25% | 95.89% |
| CHEMBL1163125 | O60885 | Bromodomain-containing protein 4 | 89.30% | 97.31% |
| CHEMBL2635 | P51452 | Dual specificity protein phosphatase 3 | 87.51% | 94.00% |
| CHEMBL3137262 | O60341 | LSD1/CoREST complex | 87.17% | 97.09% |
| CHEMBL3192 | Q9BY41 | Histone deacetylase 8 | 86.79% | 93.99% |
| CHEMBL2056 | P21728 | Dopamine D1 receptor | 86.66% | 91.00% |
| CHEMBL1293249 | Q13887 | Kruppel-like factor 5 | 86.53% | 86.33% |
| CHEMBL225 | P28335 | Serotonin 2c (5-HT2c) receptor | 86.39% | 89.62% |
| CHEMBL3880 | P07900 | Heat shock protein HSP 90-alpha | 86.28% | 96.21% |
| CHEMBL2146302 | O94925 | Glutaminase kidney isoform, mitochondrial | 85.37% | 100.00% |
| CHEMBL4208 | P20618 | Proteasome component C5 | 85.24% | 90.00% |
| CHEMBL4203 | Q9HAZ1 | Dual specificity protein kinase CLK4 | 84.94% | 94.45% |
| CHEMBL213 | P08588 | Beta-1 adrenergic receptor | 84.01% | 95.56% |
| CHEMBL1913 | P09619 | Platelet-derived growth factor receptor beta | 82.75% | 95.70% |
| CHEMBL1937 | Q92769 | Histone deacetylase 2 | 82.43% | 94.75% |
| CHEMBL3492 | P49721 | Proteasome Macropain subunit | 82.24% | 90.24% |
| CHEMBL5028 | O14672 | ADAM10 | 81.41% | 97.50% |
| CHEMBL3438 | Q05513 | Protein kinase C zeta | 80.94% | 88.48% |
| CHEMBL240 | Q12809 | HERG | 80.92% | 89.76% |
| CHEMBL3038477 | P67870 | Casein kinase II alpha/beta | 80.54% | 99.23% |
Below are displayed all the plants proven (via scientific papers) to contain this
compound!
To see more specific details click the taxa you are interested in.
To see more specific details click the taxa you are interested in.
| Ancistrocladus abbreviatus |
| Triphyophyllum peltatum |
| PubChem | 443774 |
| LOTUS | LTS0163636 |
| wikiData | Q27104194 |