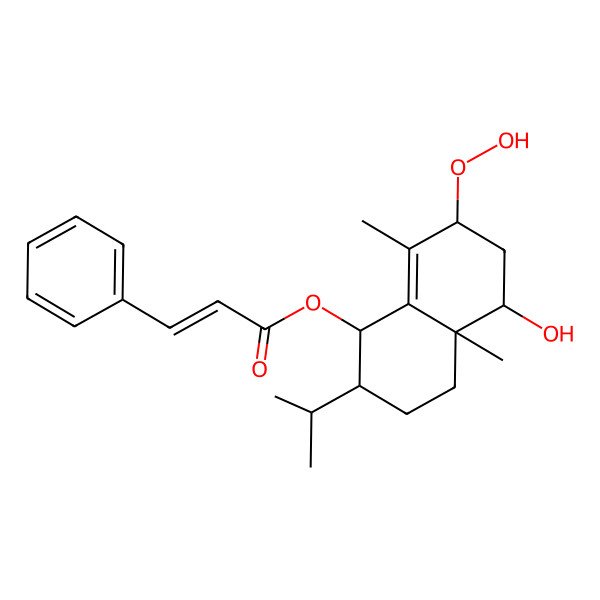
(7-hydroperoxy-5-hydroxy-4a,8-dimethyl-2-propan-2-yl-2,3,4,5,6,7-hexahydro-1H-naphthalen-1-yl) 3-phenylprop-2-enoate
| Internal ID | 7c27a08a-eac5-4377-9319-7cb0fce21056 |
| Taxonomy | Lipids and lipid-like molecules > Prenol lipids > Sesquiterpenoids > Eudesmane, isoeudesmane or cycloeudesmane sesquiterpenoids |
| IUPAC Name | (7-hydroperoxy-5-hydroxy-4a,8-dimethyl-2-propan-2-yl-2,3,4,5,6,7-hexahydro-1H-naphthalen-1-yl) 3-phenylprop-2-enoate |
| SMILES (Canonical) | CC1=C2C(C(CCC2(C(CC1OO)O)C)C(C)C)OC(=O)C=CC3=CC=CC=C3 |
| SMILES (Isomeric) | CC1=C2C(C(CCC2(C(CC1OO)O)C)C(C)C)OC(=O)C=CC3=CC=CC=C3 |
| InChI | InChI=1S/C24H32O5/c1-15(2)18-12-13-24(4)20(25)14-19(29-27)16(3)22(24)23(18)28-21(26)11-10-17-8-6-5-7-9-17/h5-11,15,18-20,23,25,27H,12-14H2,1-4H3 |
| InChI Key | RCNVBRCNVUOFOJ-UHFFFAOYSA-N |
| Popularity | 0 references in papers |
| Molecular Formula | C24H32O5 |
| Molecular Weight | 400.50 g/mol |
| Exact Mass | 400.22497412 g/mol |
| Topological Polar Surface Area (TPSA) | 76.00 Ų |
| XlogP | 3.70 |
| There are no found synonyms. |
| Target | Value | Probability (raw) | Probability (%) |
|---|---|---|---|
| No predicted properties yet! | |||
Proven Targets:
| CHEMBL ID | UniProt ID | Name | Min activity | Assay type | Source |
|---|---|---|---|---|---|
| No proven targets yet! | |||||
Predicted Targets (via Super-PRED):
| CHEMBL ID | UniProt ID | Name | Probability | Model accuracy |
|---|---|---|---|---|
| CHEMBL221 | P23219 | Cyclooxygenase-1 | 97.16% | 90.17% |
| CHEMBL2035 | P08912 | Muscarinic acetylcholine receptor M5 | 95.92% | 94.62% |
| CHEMBL3108638 | O15164 | Transcription intermediary factor 1-alpha | 95.91% | 95.56% |
| CHEMBL3251 | P19838 | Nuclear factor NF-kappa-B p105 subunit | 95.86% | 96.09% |
| CHEMBL1821 | P08173 | Muscarinic acetylcholine receptor M4 | 95.06% | 94.08% |
| CHEMBL216 | P11229 | Muscarinic acetylcholine receptor M1 | 94.05% | 94.23% |
| CHEMBL5619 | P27695 | DNA-(apurinic or apyrimidinic site) lyase | 94.04% | 91.11% |
| CHEMBL2581 | P07339 | Cathepsin D | 93.46% | 98.95% |
| CHEMBL1075094 | Q16236 | Nuclear factor erythroid 2-related factor 2 | 92.65% | 96.00% |
| CHEMBL5028 | O14672 | ADAM10 | 90.94% | 97.50% |
| CHEMBL3137262 | O60341 | LSD1/CoREST complex | 89.21% | 97.09% |
| CHEMBL3091268 | Q92753 | Nuclear receptor ROR-beta | 88.14% | 95.50% |
| CHEMBL245 | P20309 | Muscarinic acetylcholine receptor M3 | 88.04% | 97.53% |
| CHEMBL1293249 | Q13887 | Kruppel-like factor 5 | 87.81% | 86.33% |
| CHEMBL4793 | Q86TI2 | Dipeptidyl peptidase IX | 87.45% | 96.95% |
| CHEMBL5608 | Q16288 | NT-3 growth factor receptor | 84.60% | 95.89% |
| CHEMBL3130 | O00329 | PI3-kinase p110-delta subunit | 84.28% | 96.47% |
| CHEMBL3359 | P21462 | Formyl peptide receptor 1 | 83.79% | 93.56% |
| CHEMBL2716 | Q8WUI4 | Histone deacetylase 7 | 83.65% | 89.44% |
| CHEMBL3807 | P17706 | T-cell protein-tyrosine phosphatase | 83.20% | 93.00% |
| CHEMBL211 | P08172 | Muscarinic acetylcholine receptor M2 | 82.07% | 94.97% |
| CHEMBL1806 | P11388 | DNA topoisomerase II alpha | 81.84% | 89.00% |
| CHEMBL4227 | P25090 | Lipoxin A4 receptor | 81.51% | 100.00% |
| PubChem | 162864920 |
| LOTUS | LTS0063650 |
| wikiData | Q105233827 |